Nanotether Technology
The company’s unique technology will transform the identification and characterisation of biomolecular interactions. Quantitative analysis of such interactions is critical across all areas of the biological sciences, particularly drug discovery where the race for new compounds and formulations is intensely competitive.
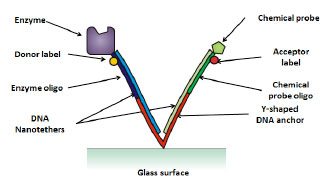
The Nanotether system, depicted in figure 1, illustrates the interaction between a protein enzyme and chemical probe, which are both are attached to the ends of nanoscale tethers, which are in turn bound to a glass surface. Both tethers and the anchor molecule are composed of DNA. The enzyme and chemical probe are free to interact and reversibly bind. The degree to which this binding is disrupted in the presence of a competing chemical compound provides information on whether the chemical entity is a potential drug for this target.
Figure 1
Basic structure of a Nanotether consisting of a protein enzyme and a chemical probe which are known to bind reversibly.
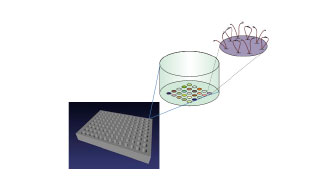
Features containing thousands of Nanotether molecules are printed in spots onto a solid surface (figure 2), with each feature being composed of a different population of paired biomolecules that are free to interact. Nanotether features are typically 100μm to 3mm in diameter, depending on the application and format.
Figure 2
Nanotethers are arrayed out as features (or spots) in a grid pattern on the bottom of microtitre plate wells. Each well can be filled with different biological or chemical components to observe and measure biomolecular interactions.
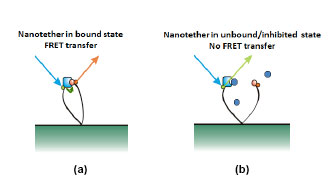
Potential drug compounds then flow into the system to disrupt the bound protein-probe pair. Nanotethers can be observed in one of two states: bound and unbound. In the bound state (figure 3a) the chemical probe docks into the binding site of the protein enzyme. In this state the incoming light pulse is absorbed by the protein (donor) label, undergoes energy transfer to the probe (acceptor) label and is then re-emitted at a different wavelength. In the unbound state (figure 3b) a potential drug molecule has docked into the binding site of the protein enzyme blocking the probe from binding. In this scenario the protein (donor) label absorbs the laser pulse, but with probe (acceptor) label not in close proximity, there is no energy transfer and the pulse is re-emitted from the donor label. As the pulses from the two labels are emitted at different wavelengths it is straightforward to measure the ratio, and hence the relative proportions of the bound and unbound Nanotether states for each feature/assay.
Figure 3
Nanotethers are either in a bound (a) or unbound state (b). In (b) a potential drug molecule (blue sphere) is blocking the enzymes binding site. The incoming light pulses (blue arrow) are re-emitted back at different wavelengths indicating whether the Nanotether is in the bound (orange arrow) or unbound (green arrow) state. The ratio of Nanotethers in the bound to unbound state provides an indication of the potential of the chemical compounds being a suitable drug.
Publications
- Kasry, A., Borri, P., Davies, P. R., Harwood, A., Thomas, N., Lofas, S., & Dale, T. (2009). Comparison of methods for generating planar DNA-modified surfaces for hybridization studies. ACS applied materials & interfaces, ACS applied materials & interfaces, 1(8), 1793–1798.
- Perrins, R. D., Orchard, C., Zavodszky, M., Kasry, A., Nikolaev, N., Harwood, A., Borri, P., et al. (2011). Doing more with less: a method for low total mass, affinity measurement using variable-length nanotethers Anal Chem, 111026113737007. doi:10.1021/ac2012569.
- Doing more with less: low mass affinity measurement using variable-length nanotethers (2013). Richard Perrins, Gareth Bray, Dan Leggate, Bei Li, Meike Roskamp, Markos Trikeriotis and Marcus Harrison. Nanotether Discovery Sciences Ltd, The Sir Martin Evans Building, Museum Avenue, Cardiff, CF10 3AX, UK. Paola Borri, Adrian Harwood and Trevor Dale. Cardiff School of Biosciences, Biomedical Sciences Building, Museum Avenue, Cardiff, CF10 3AX, UK